
Polypurine reverse-Hoogsteen hairpins (PPRHs) are non-modified oligonucleotides containing two polypurine domains, in a mirror repeat fashion, linked by a pentathymidine stretch forming double-stranded DNA stem-loop molecules. The two polypurine domains interact by intramolecular reverse-Hoogsteen bonds allowing the formation of this specific hairpin structure.
Properties

PPRHs can bind to polypyrimidine stretches in either single- or double stranded DNA by Watson and Crick bonds establishing triple-stranded DNA structures. The formation of PPRHs triplexes takes place at physiological pH. PPRHs provoke a strand displacement.[1] of the homopurine sequence of the target dsDNA, opening the two strands of the DNA. There are two types of PPRHs: i) Template-PPRHs[2] that bind to the template strand of DNA, inhibiting transcription; and ii) Coding-PPRHs[3] that bind to the coding strand of the DNA altering splicing. Both types of PPRHs decrease gene expression. PPRHs present high stability in serum and cells and show lack of immunogenicity not activating the innate inflammatory response.[4] PPRHs do not have off-target effects and do not show hepatotoxicity or nephrotoxicity.[5]
Applications
PPRHs could be used as gene silencing tools[6] acting by different mechanisms than triplex forming oligonucleotides (TFOs), antisense oligonucleotides or siRNAs. Upon binding to their targets, PPRHs can decrease the mRNA and protein levels of the selected genes. Their action has been demonstrated in vitro for a number of genes involved in metabolism (DHFR), proliferation (mTOR), DNA topology (TOP1), lifespan and senescence (telomerase), apoptosis (survivin, BCL2), transcription factors and non-druggable targets (c-MYC[7] and k-Ras[8]) , proto-oncogenes (MDM2),[9] replication stress (WEE1, CHK1)[10] and Thymidilate synthase (TYMS)[11] as part of a cancer gene therapy strategy. Their preclinical proof of principle has been proven in vivo using the antiapoptotic survivin gene.[12] PPRHs have also been applied as tools in cancer immunotherapy by silencing CD47 in MCF7 breast cancer cells and SIRPα in macrophages,[13] and the PD-1/PD-L1 pathway in human tumor cells.[14][15] PPRHs can also be used as the capture probe in different devises to detect viral infection by forming a triplex with the RNA of the virus such as SARS-CoV-2 in a technology termed Triplex Enhanced Nucleic Acid Detection Assay (TENADA)[16]
Design and improvements

PPRHs can be designed for virtually any gene in the genome by searching for polypirimidine stretches in the sequence of the desired gene. Optimal lengths for each domain of the PPRHs are within 20–30 nucleotides. The total length of a typical PPRH is 55 nucleotides considering two domains of 25 bases plus 5T for the linking loop. If purine interruptions are encountered (up to three) within the polypirimidine target, the highest affinity of PPRH binding is achieved by placing in the hairpin the complementary base (a pyrimidine) in front of the purines[17] (Wild type-PPRH).
Wedge-PPRH
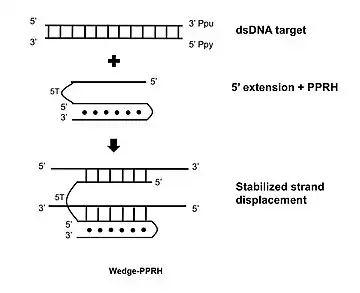
A further development consists in extending the 5' flank of the PPRH with a sequence complementary to the displaced polypurine strand of the target dsDNA which stabilizes the strand displacement, producing additional binding and functionality.[17]
WEB tools
A triplex target DNA site (TTS), a stretch of DNA that is composed of polypurines, is able to form a triple-helix (triplex) structure in genomic DNA. Integrative WEB tools for identification and analysis of the triplex formation target DNA sequences, including PPRH sequences, associated with genes and regulatory elements (e.g., transcription factor binding sites, repeats, G-quadruplet motifs, SNPs, and non-protein coding regulatory DNA elements) in the human genome are publicly available (see External links).[18][19]
These tools could be used to search biologically meaningful genome polypurine stretches, help to understand biological roles of the natural paired polypurine domains like PPRH and to optimize experimental design of anti-gene treatment.
References
- ↑ Coma S, Noé V, Eritja R, Ciudad CJ (December 2005). "Strand displacement of double-stranded DNA by triplex-forming antiparallel purine-hairpins". Oligonucleotides. 15 (4): 269–283. doi:10.1089/oli.2005.15.269. hdl:10261/124878. PMID 16396621.
- ↑ de Almagro MC, Coma S, Noé V, Ciudad CJ (April 2009). "Polypurine hairpins directed against the template strand of DNA knock down the expression of mammalian genes". The Journal of Biological Chemistry. 284 (17): 11579–11589. doi:10.1074/jbc.M900981200. PMC 2670163. PMID 19261618.
- ↑ de Almagro MC, Mencia N, Noé V, Ciudad CJ (April 2011). "Coding polypurine hairpins cause target-induced cell death in breast cancer cells". Human Gene Therapy. 22 (4): 451–463. doi:10.1089/hum.2010.102. PMID 20942657.
- ↑ Villalobos X, Rodríguez L, Prévot J, Oleaga C, Ciudad CJ, Noé V (January 2014). "Stability and immunogenicity properties of the gene-silencing polypurine reverse Hoogsteen hairpins". Molecular Pharmaceutics. 11 (1): 254–264. doi:10.1021/mp400431f. PMID 24251728.
- ↑ Félix AJ, Ciudad CJ, Noé V (September 2018). "Functional pharmacogenomics and toxicity of PolyPurine Reverse Hoogsteen hairpins directed against survivin in human cells". Biochemical Pharmacology. 155: 8–20. doi:10.1016/j.bcp.2018.06.020. PMID 29940174. S2CID 49405355.
- ↑ Aviñó A, Eritja R, Ciudad CJ, Noé V (June 2019). "Parallel Clamps and Polypurine Hairpins (PPRH) for Gene Silencing and Triplex-Affinity Capture: Design, Synthesis, and Use". Current Protocols in Nucleic Acid Chemistry. 77 (1): e78. doi:10.1002/cpnc.78. hdl:10261/182909. PMID 30912630. S2CID 85517931.
- ↑ Valiuska S, Psaras AM, Noé V, Brooks TA, Ciudad CJ (December 2022). "Targeting MYC Regulation with Polypurine Reverse Hoogsteen Oligonucleotides". International Journal of Molecular Sciences. 24 (1): 378. doi:10.3390/ijms24010378. PMC 9820101. PMID 36613820.
- ↑ Psaras AM, Valiuska S, Noé V, Ciudad CJ, Brooks TA (February 2022). "Targeting KRAS Regulation with PolyPurine Reverse Hoogsteen Oligonucleotides". International Journal of Molecular Sciences. 23 (4): 2097. doi:10.3390/ijms23042097. PMC 8876201. PMID 35216221.
- ↑ Villalobos X, Rodríguez L, Solé A, Lliberós C, Mencia N, Ciudad CJ, Noé V (August 2015). "Effect of Polypurine Reverse Hoogsteen Hairpins on Relevant Cancer Target Genes in Different Human Cell Lines". Nucleic Acid Therapeutics. 25 (4): 198–208. doi:10.1089/nat.2015.0531. PMID 26042602.
- ↑ Aubets E, Noé V, Ciudad CJ (May 2020). "Targeting replication stress response using polypurine reverse hoogsteen hairpins directed against WEE1 and CHK1 genes in human cancer cells". Biochemical Pharmacology. 175: 113911. doi:10.1016/j.bcp.2020.113911. PMID 32173365. S2CID 212728120.
- ↑ Aubets E, J Félix A, Garavís M, Reyes L, Aviñó A, Eritja R, et al. (July 2020). "Detection of a G-Quadruplex as a Regulatory Element in Thymidylate synthase for Gene Silencing Using Polypurine Reverse Hoogsteen Hairpins". International Journal of Molecular Sciences. 21 (14): 5028. doi:10.3390/ijms21145028. PMC 7404261. PMID 32708710.
- ↑ Rodríguez L, Villalobos X, Dakhel S, Padilla L, Hervas R, Hernández JL, et al. (December 2013). "Polypurine reverse Hoogsteen hairpins as a gene therapy tool against survivin in human prostate cancer PC3 cells in vitro and in vivo". Biochemical Pharmacology. 86 (11): 1541–1554. doi:10.1016/j.bcp.2013.09.013. PMID 24070653.
- ↑ Bener G, J Félix A, Sánchez de Diego C, Pascual Fabregat I, Ciudad CJ, Noé V (September 2016). "Silencing of CD47 and SIRPα by Polypurine reverse Hoogsteen hairpins to promote MCF-7 breast cancer cells death by PMA-differentiated THP-1 cells". BMC Immunology. 17 (1): 32. doi:10.1186/s12865-016-0170-z. PMC 5037635. PMID 27671753.
- ↑ Medina Enríquez MM, Félix AJ, Ciudad CJ, Noé V (2018). "Cancer immunotherapy using PolyPurine Reverse Hoogsteen hairpins targeting the PD-1/PD-L1 pathway in human tumor cells". PLOS ONE. 13 (11): e0206818. Bibcode:2018PLoSO..1306818M. doi:10.1371/journal.pone.0206818. PMC 6219785. PMID 30399174.
- ↑ Ciudad CJ, Medina Enriquez MM, Félix AJ, Bener G, Noé V (April 2019). "Silencing PD-1 and PD-L1: the potential of PolyPurine Reverse Hoogsteen hairpins for the elimination of tumor cells". Immunotherapy. 11 (5): 369–372. doi:10.2217/imt-2018-0215. PMID 30786843. S2CID 73467979.
- ↑ Aviñó A, Cuestas-Ayllón C, Gutiérrez-Capitán M, Vilaplana L, Grazu V, Noé V, et al. (December 2022). "Detection of SARS-CoV-2 Virus by Triplex Enhanced Nucleic Acid Detection Assay (TENADA)". International Journal of Molecular Sciences. 23 (23): 15258. doi:10.3390/ijms232315258. PMC 9740288. PMID 36499587.
- 1 2 Rodríguez L, Villalobos X, Solé A, Lliberós C, Ciudad CJ, Noé V (March 2015). "Improved design of PPRHs for gene silencing". Molecular Pharmaceutics. 12 (3): 867–877. doi:10.1021/mp5007008. PMID 25615267.
- ↑ Jenjaroenpun P, Kuznetsov VA (December 2009). "TTS mapping: integrative WEB tool for analysis of triplex formation target DNA sequences, G-quadruplets and non-protein coding regulatory DNA elements in the human genome". BMC Genomics. 10 (Suppl 3): S9. doi:10.1186/1471-2164-10-S3-S9. PMC 2788396. PMID 19958507.
- ↑ Jenjaroenpun P, Chew CS, Yong TP, Choowongkomon K, Thammasorn W, Kuznetsov VA (January 2015). "The TTSMI database: a catalog of triplex target DNA sites associated with genes and regulatory elements in the human genome". Nucleic Acids Research. 43 (Database issue): D110–D116. doi:10.1093/nar/gku970. PMC 4384029. PMID 25324314.
External links
- "Triplex-Forming Oligonucleotide Target Sequence Search Tool". The University of Texas MD Anderson Cancer Center. A Searching Tool to find Polypurine and Polypyrimidine stretches in DNA
- Jenjaroenpun P, Chew CS, Yong TP, Choowongkomon K, Thammasorn W, Kuznetsov VA. "The TTSMI database: a catalog of triplex target DNA sites associated with genes and regulatory elements in the human genome". A*STAR Biomedical Sciences Institutes.
- Jenjaroenpun P, Kuznetsov VA. "TTS mapping: integrative WEB tool for analysis of triplex formation target DNA sequences, G-quadruplets and non-protein coding regulatory DNA elements in the human genome". A*STAR Biomedical Sciences Institutes.